Lipoprotein(a) Genomics
About
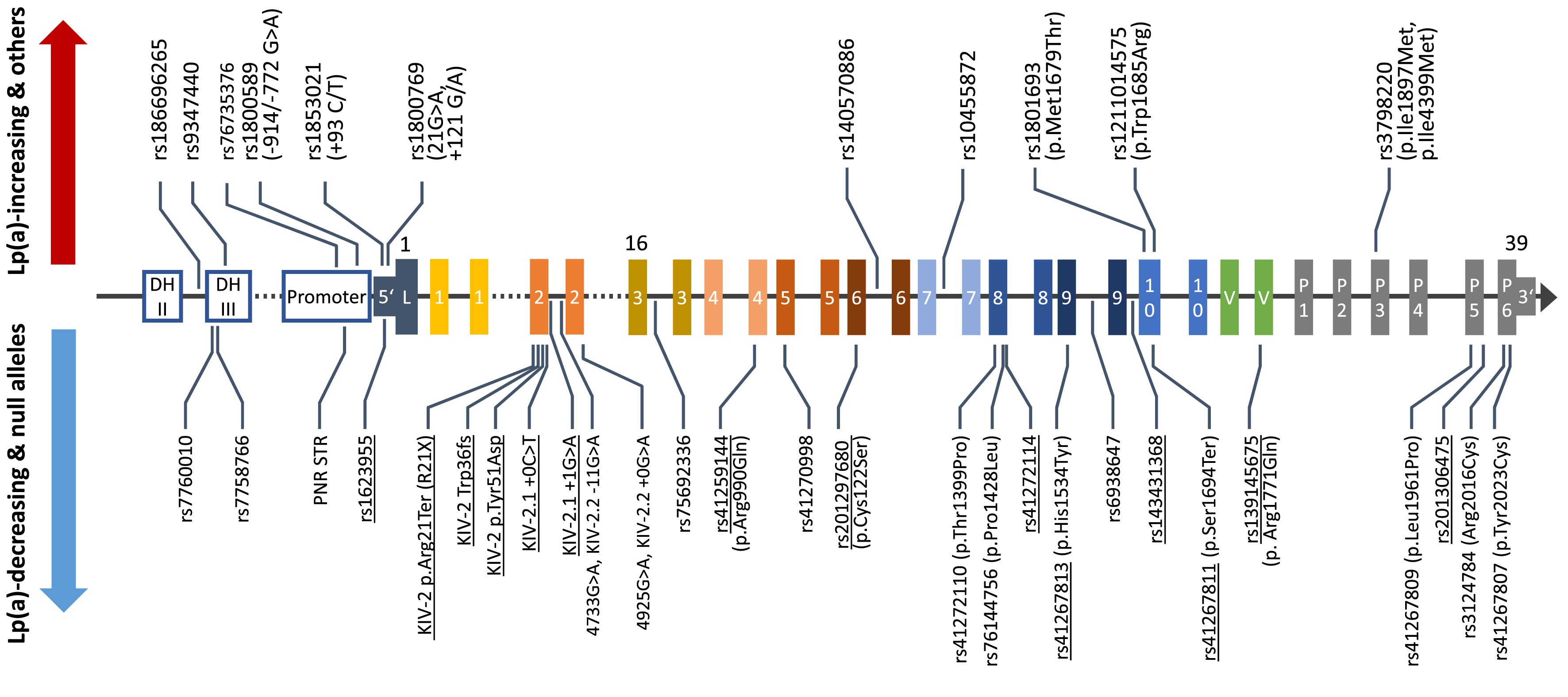
Genetic variation in the LPA gene explains up to 90% of lipoprotein(a) [Lp(a)] plasma concentrations in the population but the genetic mechanisms that govern the individual lipoprotein(a) concentrations are poorly understood. We investigate how functional genetic variants in LPA create the huge variance in lipoprotein(a) concentrations that we observe between individuals and across ancestries. A major part of our work concerns the investigation of mutations located in the KIV-2 repeat region of the LPA gene. This repetitive region can encompass up to 70% of the coding sequence of the gene, but, due to its complex structure, it has not been accessible until recently. By combining the computational expertise of the institute in the detection of low-level mutations, advanced molecular techniques genetics and our long-standing experience in Lp(a) research, we have developed a comprehensive genetic toolset capable of adressing many difficult aspects of Lp(a) genetics. Additionally, we enjoy exploring the potential of nanopore sequencing and other emerging genetic technologies to tackle tough genetic questions across disciplines.
Team

Professor of Computational Genomics
+43 512 9003 70579
sebastian.schoenherr@i-med.ac.at

Lab technician (BMA) (on maternal leave)
+43 512 9003 70573
gertraud.streiter@i-med.ac.at
Technologies available to dissect the complex genetics of lipoprotein(a)
- A comprehensive toolbox of advanced PCR technologies
- Long-read sequencing (Nanopore sequencing, as well as PacBio SMRT sequencing via cooperations)
- KIV-2 variant detection by Next-generation sequencing (NGS)
- Ultrasenstive KIV-2 mutation typing by droplet digital PCR (ddPCR), allele-specific qPCR and castPCR
- Roboter-assisted high-throughput sample preparation, PCR/qPCR pipetting and TaqMan genotyping
- Cloning techniques, minigene splicing assays and luciferase reporter assays
- Ultra-precise haplotyping of variants in the LPA KIV-2 VNTR by UMI-corrected nanopore sequencing
- Pulsed-field gel electrophoresis (PFGE) (LPA allele sizing and mutation phasing, HMW DNA assessment)
- KIV-2 copy number determination by qPCR and droplet digital PCR
- HepG2 cell lines expressing various apolipoprotein(a) isoforms
Key Publications
Coassin S, Kronenberg F: Lipoprotein(a) beyond the kringle IV repeat polymorphism: The complexity of genetic variation in the LPA gene. Atherosclerosis 349:17-35, 2022. PMID: 35606073 Review
Amstler S, Streiter G, Pfurtscheller C, Forer L, Di Maio S, Weissensteiner H, Paulweber B, Schönherr S, Kronenberg F, Coassin S: Nanopore sequencing with unique molecular identifiers enables accurate mutation analysis and haplotyping in the complex lipoprotein(a) KIV-2 VNTR. Genome Med. 16:117, 2024. PMID: 39380090 Journal Article
Grüneis R, Weissensteiner H, Lamina C, Schönherr S, Forer L, Di Maio S, Streiter G, Peters A, Gieger C, Kronenberg F, Coassin S: The kringle IV type 2 domain variant 4925G>A causes the elusive association signal of the LPA pentanucleotide repeat. J. Lipid Res. 63:100306, 2022. PMID: 36309064 Journal Article
Grüneis R, Lamina C, Di Maio S, Schönherr S, Zoescher P, Forer L, Streiter G, Peters A, Gieger C, Köttgen A, Kronenberg F, Coassin S: The effect of LPA Thr3888Pro on lipoprotein(a) and coronary artery disease is modified by the LPA KIV-2 variant 4925G>A. Atherosclerosis 349:151-159, 2022. PMID: 35534298 Journal Article
Schachtl-Riess JF, Kheirkhah A, Grüneis R, Di Maio S, Schoenherr S, Streiter G, Losso JL, Paulweber B, Eckardt KU, Köttgen A, Lamina C, Kronenberg F, Coassin S, GCKD Investigators: Frequent LPA KIV-2 variants lower lipoprotein(a) concentrations and protect against coronary artery disease. J. Am. Coll. Cardiol. 78:437-449, 2021. PMID: 34325833 Journal Article
Di Maio S, Grüneis R, Streiter G, Lamina C, Maglione M, Schoenherr S, Öfner D, Thorand B, Peters A, Eckardt KU, Köttgen A, Kronenberg F, Coassin S: Investigation of a nonsense mutation located in the complex KIV-2 copy number variation region of apolipoprotein(a) in 10,910 individuals. Genome Med. 12:74, 2020. PMID: 32825847 Journal Article
Coassin S, Schönherr S, Weissensteiner H, Erhart G, Forer L, Losso JL, Lamina C, Haun M, Utermann G, Paulweber B, Specht G, Kronenberg F: A comprehensive map of single-base polymorphisms in the hypervariable LPA kringle IV type 2 copy number variation region. J. Lipid Res. 60:186-199, 2019. PMID: 30413653 Journal Article
Coassin S, Erhart G, Weissensteiner H, Eca Guimarães de Araújo M, Lamina C, Schönherr S, Forer L, Haun M, Losso JL, Köttgen A, Schmidt K, Utermann G, Peters A, Gieger C, Strauch K, Finkenstedt A, Bale R, Zoller H, Paulweber B, Eckardt KU, Hüttenhofer A, Huber LA, Kronenberg F: A novel but frequent variant in LPA KIV-2 is associated with a pronounced Lp(a) and cardiovascular risk reduction. Eur. Heart J. 38:1823-1831, 2017. PMID: 28444229 Journal Article




